Using Loops In R
This document is a practical guide for getting started using for and while loops in R. My goal is to help readers who are familiar with R, but unfamiliar with using loops in R, get started coding with loops. This guide was original written for an event in Stirling Coding Club, and a version of this document (formatted slightly differently) is available here.
- Introduction: What is a loop?
- The for loop in R: getting started
- The for loop in R: a real example
- The for loop in R: nested loops
- The while loop in R
- Practice problems
- Additional resources
Introduction: What is a loop?
Being able to use loops is a critical skill in programming and working with large arrays of data. Loops make it possible to repeat a set of instructions (i.e., code) for a particular set of conditions (e.g., for a range of numbers from 1 to 1000), or while a set of conditions still applies (e.g., while a value is still greater than zero). Hence, the use of for loops and while loops are fundamental for writing and running code efficiently (note that other types of loops also exist, but I will not focus on them now). Here I will introduce the key concepts of programming with loops, with particular emphasis on getting started with some practical uses of loops in the R programming language for scientific researchers.
The R programming language includes many base level functions to perform tasks that would otherwise require loops (e.g., functions such as apply and tapply, which effectively repeat a set of instructions to summarise values in an array). Tens of thousands of downloadable packages (code, data, and documentation bundled together, written by and for R users) are available in R, most of which include their own functions that can perform specific tasks required in scientific research (e.g., vegan, dplyr, and shiny). Hence, successful data analysis in R can often just be a matter of finding and using an appropriate and reliable package for a given task. Nevertheless, unique data sets and models often require unique code, so base and package functions cannot always be found to do custom tasks. In many situations, the ability to use loops to repeat tasks will make it possible to quickly and confidently develop reproducible code in data analysis. In the long term, this will likely save time; in the short term, it creates an opportunity to develop and practice coding skills.
Below I will demonstrate how to use for loops and while loops.
- A for loop iterates a set of instructions for a predetermined set of conditions (i.e., when you know how many times you need to iterate the same set of instructions)
- A while loop iterates a set of instructions while some condition(s) remains satisfied (i.e., when you might not know how many times you need to iterate the same set of instructions, but you do know when the iterations need to stop)
It is not important to completely understand the two definitions above before getting started, just as it is not important to understand a verbal definition of ‘multiplication’ before learning to multiply two numbers. Seeing examples of loops in practice will make it clear how they work and when to use them. In the next section, I will therefore start with some key examples to show how for loops can be used in R. I will then do the same with while loops. Finally, I will provide some practice problems and additional resources for using loops in programming.
The for loop in R: getting started
Different languages have differ syntaxes for writing for loops. In R, the syntax is as follows:
for(index in set_of_conditions){
# Code that gets repeated for each condition
}
In the above, anything within the bracketed { } gets repeated with index being substituted, sequentially, for all possible conditions in the set_of_conditions. A more concrete example will help:
for(i in 1:10){
print(i);
}
We can interpret the line for(i in 1:10) verbally to explain what is going on in plain English:
Substitute the index
ifor each value from 1 to 10 (i.e., 1, 2, 3, …, 8, 9, 10), and run the following bracketed code with each value in sequence. In other words, run the bracketed code first withi = 1, then withi = 2, and so forth until finishing the loop withi = 10.
Given the above explanation, we can predict what will happen with the example code above, which I now run below.
for(i in 1:10){
print(i);
}
## [1] 1
## [1] 2
## [1] 3
## [1] 4
## [1] 5
## [1] 6
## [1] 7
## [1] 8
## [1] 9
## [1] 10
The code has printed integers from 1 to 10. This is obviously a very simple example of using a loop, but it highlights the basic idea. There are three key points that I want to note with this example before moving on
1. the above for loop is effectively doing the same as the code below
print(1);
print(2);
print(3);
print(4);
print(5);
print(6);
print(7);
print(8);
print(9);
print(10);
## [1] 1
## [1] 2
## [1] 3
## [1] 4
## [1] 5
## [1] 6
## [1] 7
## [1] 8
## [1] 9
## [1] 10
In the above, I have unrolled the for loop that printed off integers from 1 to 10. But the result is the same. The advantage of using the loop is that it avoids the need to repeat the same code more times than is necessary (consider if we wanted to print values from 1 to 10000 – much less needs to be changed when using the for loop, and much fewer lines of code need to be written). There is no hard rule for when to use a loop versus when to repeat the same line(s) of code multiple times; it is generally best to use whichever method is most readable to (future) you. In practice, when getting started, it might help to think about what the loops would look like if unrolled – or even write them both ways to confirm that a loop is working as intended.
2. There is nothing special about i
In a lot of books and online examples introducing loops, the index i is used as it is in my above example. But there is nothing special about i, just as there is nothing special about the variable x in algebra. The index i just serves as a placeholder for whatever actual value is going to be substituted from the set of conditions (i.e., values from 1 to 10 in the above example). We get the exact same result with the following code:
for(value_to_be_printed in 1:10){
print(value_to_be_printed);
}
## [1] 1
## [1] 2
## [1] 3
## [1] 4
## [1] 5
## [1] 6
## [1] 7
## [1] 8
## [1] 9
## [1] 10
The above use of the long value_to_be_printed instead of the shorter i seems unnecessary at first, but it is actually often helpful to give indices specific names like this to make code more readable. Doing so becomes especially helpful when working with multiple indices and loops within loops (an example of this later), or when the number of lines between the starting { and ending } brackets becomes larger than can be viewed on a computer screen.
3. There is nothing special about 1:10
A lot of introductions to for loops in R will show the set of values to be iterated in this way, but there are equally acceptable ways to write it. For example, consider the below:
for(i in c(1, 2, 3, 4, 5, 6, 7, 8, 9, 10)){
print(i);
}
## [1] 1
## [1] 2
## [1] 3
## [1] 4
## [1] 5
## [1] 6
## [1] 7
## [1] 8
## [1] 9
## [1] 10
Or even the below, where I first define the set of values with its own variable named the_set:
the_set <- c(1, 2, 3, 4, 5, 6, 7, 8, 9, 10);
for(i in the_set){
print(i);
}
## [1] 1
## [1] 2
## [1] 3
## [1] 4
## [1] 5
## [1] 6
## [1] 7
## [1] 8
## [1] 9
## [1] 10
The order of numbers does matter. For example, we could reverse the order in which numbers are printed by reversing the set of values:
for(i in 10:1){
print(i);
}
## [1] 10
## [1] 9
## [1] 8
## [1] 7
## [1] 6
## [1] 5
## [1] 4
## [1] 3
## [1] 2
## [1] 1
We could even print off the numbers in a random order with the below:
random_vec <- sample(x = 1:10, size = 10);
for(i in random_vec){
print(i);
}
## [1] 3
## [1] 6
## [1] 10
## [1] 4
## [1] 2
## [1] 9
## [1] 1
## [1] 5
## [1] 7
## [1] 8
It is unlikely that there would ever be a need to reverse the order of a set, and for most for loops, the simple 1:N format will usually be all that that is needed. The point is that there is no reason to feel constrained to using this format when writing loops.
The for loop in R: a real example
The examples above were intended only to introduce the general idea of using for loops, and their specific syntax in R. In coding, there is rarely such a need to use for loops to simply print off values. More often, for loops are used to repeat the same set of (often complex) instructions for a set of values. As a real example, I will use use the nhtemp data set in R.
data(nhtemp); # Reads in the data set
The nhtemp data set includes a vector that stores the mean annual temperature in degrees Fahrenheit in New Haven, Connecticut (USA), from 1912 to 1971.
print(nhtemp);
## Time Series:
## Start = 1912
## End = 1971
## Frequency = 1
## [1] 49.9 52.3 49.4 51.1 49.4 47.9 49.8 50.9 49.3 51.9 50.8 49.6 49.3 50.6
## [15] 48.4 50.7 50.9 50.6 51.5 52.8 51.8 51.1 49.8 50.2 50.4 51.6 51.8 50.9
## [29] 48.8 51.7 51.0 50.6 51.7 51.5 52.1 51.3 51.0 54.0 51.4 52.7 53.1 54.6
## [43] 52.0 52.0 50.9 52.6 50.2 52.6 51.6 51.9 50.5 50.9 51.7 51.4 51.7 50.8
## [57] 51.9 51.8 51.9 53.0
In the above data, the first value 49.9 is therefore the mean temperature from 1912, and the last value 53 is the mean temperature from 1971. We can print these off by using the indices nhtemp[1] and nhtemp[60] (as there are length(nhtemp) = 60 temperature years in nhtemp).
print(nhtemp[1]);
## [1] 49.9
print(nhtemp[60]);
## [1] 53
We might want to use these data to analyse how the temperature in New Haven has changed over the years from 1912-1972. The first task would likely be to convert the temperatures from Fahrenheit to Celsius. The formula for conversion is as follows,
To get , it is not actually necessary to use a for loop in R; this can be done in one line of code:
T_Celsius <- (5/9) * (nhtemp - 32);
print(T_Celsius);
## Time Series:
## Start = 1912
## End = 1971
## Frequency = 1
## [1] 9.944444 11.277778 9.666667 10.611111 9.666667 8.833333 9.888889
## [8] 10.500000 9.611111 11.055556 10.444444 9.777778 9.611111 10.333333
## [15] 9.111111 10.388889 10.500000 10.333333 10.833333 11.555556 11.000000
## [22] 10.611111 9.888889 10.111111 10.222222 10.888889 11.000000 10.500000
## [29] 9.333333 10.944444 10.555556 10.333333 10.944444 10.833333 11.166667
## [36] 10.722222 10.555556 12.222222 10.777778 11.500000 11.722222 12.555556
## [43] 11.111111 11.111111 10.500000 11.444444 10.111111 11.444444 10.888889
## [50] 11.055556 10.277778 10.500000 10.944444 10.777778 10.944444 10.444444
## [57] 11.055556 11.000000 11.055556 11.666667
But if we know in advance that we will be doing some more complex data manipulation later, in might make sense to start out doing the conversion within a loop instead. To use a loop, we can first define the vector T_Celsius, then apply the conversion for each value in nhtemp.
T_Celsius <- NULL;
for(year in 1:length(nhtemp)){
T_Celsius[year] <- (5/9) * (nhtemp[year] - 32);
}
print(T_Celsius);
## [1] 9.944444 11.277778 9.666667 10.611111 9.666667 8.833333 9.888889
## [8] 10.500000 9.611111 11.055556 10.444444 9.777778 9.611111 10.333333
## [15] 9.111111 10.388889 10.500000 10.333333 10.833333 11.555556 11.000000
## [22] 10.611111 9.888889 10.111111 10.222222 10.888889 11.000000 10.500000
## [29] 9.333333 10.944444 10.555556 10.333333 10.944444 10.833333 11.166667
## [36] 10.722222 10.555556 12.222222 10.777778 11.500000 11.722222 12.555556
## [43] 11.111111 11.111111 10.500000 11.444444 10.111111 11.444444 10.888889
## [50] 11.055556 10.277778 10.500000 10.944444 10.777778 10.944444 10.444444
## [57] 11.055556 11.000000 11.055556 11.666667
There are a few things to note here.
- In the first line of the above, using the
repfunction, I have definedT_Celsiusto be a null vector. - I have used the index
yearrather thanito make it easier to remember what I am looping over. - The loop goes from 1 to the length of
nhtemp(length(nhtemp)= 60). I could have instead just written1:60, but the above has the advantage that if the length ofnhtempchanges for some reason, the loop will still work. As a exercise, try running the above code for1:40or1:80to see what happens when the number of years to loop over does not match the number of years innhtemp.
Now say that we actually want to know the change in temperature from one year to the next, and to make a new vector Temp_ch that stores the difference in degrees Celsius from year to year - 1. While there are ways to make such a vector in R without a loop, using a for loop is probably most intuitive way to do it.
T_Celsius <- NULL;
Temp_ch <- NULL;
for(year in 1:length(nhtemp)){
T_Celsius[year] <- (5/9) * (nhtemp[year] - 32);
Temp_ch[year] <- T_Celsius[year] - T_Celsius[year - 1];
}
print(T_Celsius);
## [1] 9.944444 11.277778 9.666667 10.611111 9.666667 8.833333 9.888889
## [8] 10.500000 9.611111 11.055556 10.444444 9.777778 9.611111 10.333333
## [15] 9.111111 10.388889 10.500000 10.333333 10.833333 11.555556 11.000000
## [22] 10.611111 9.888889 10.111111 10.222222 10.888889 11.000000 10.500000
## [29] 9.333333 10.944444 10.555556 10.333333 10.944444 10.833333 11.166667
## [36] 10.722222 10.555556 12.222222 10.777778 11.500000 11.722222 12.555556
## [43] 11.111111 11.111111 10.500000 11.444444 10.111111 11.444444 10.888889
## [50] 11.055556 10.277778 10.500000 10.944444 10.777778 10.944444 10.444444
## [57] 11.055556 11.000000 11.055556 11.666667
In the new line of code added within the above loop, the temperature in degrees Celsius from the previous year T_Celsius[year - 1] is subtracted from the tempurature from the current year T_Celsius[year]. The value of this difference is then stored in Temp_ch[year], so at the end of the loop, each element of Temp_ch stores the difference between the current year’s temperature and the last year’s temperature.
Note that this newly added line within the for loop is a bit dangerous because T_Celsius[year - 1] does not exist when year = 1 (i.e., at the start of the loop). Since there is no value at T_Celsius[1 - 1], R returns an NA, so the first value of Temp_ch = NA. This is what we want, but if we are not careful, trusting R to fill things in appropriately might cause us problems later. It is usually better to err on the side of caution and think carefully about what each line is doing, using comments to help the readability. The example below makes everything a bit cleaner and clearer.
total_years <- length(nhtemp); # Total years in the data set
# Make vectors with an NA element for each year
T_Celsius <- rep(x = NA, times = total_years);
Temp_ch <- rep(x = NA, times = total_years);
for(year in 1:total_years){ # For each year in the data set
# First calculate the temperature in degrees Celsius
T_Celsius[year] <- (5/9) * (nhtemp[year] - 32);
if(year > 1){ # Condition in which difference exists
Temp_ch[year] <- T_Celsius[year] - T_Celsius[year - 1];
} # Now T_Celsius[0] will not be attempted
}
print(T_Celsius);
## [1] 9.944444 11.277778 9.666667 10.611111 9.666667 8.833333 9.888889
## [8] 10.500000 9.611111 11.055556 10.444444 9.777778 9.611111 10.333333
## [15] 9.111111 10.388889 10.500000 10.333333 10.833333 11.555556 11.000000
## [22] 10.611111 9.888889 10.111111 10.222222 10.888889 11.000000 10.500000
## [29] 9.333333 10.944444 10.555556 10.333333 10.944444 10.833333 11.166667
## [36] 10.722222 10.555556 12.222222 10.777778 11.500000 11.722222 12.555556
## [43] 11.111111 11.111111 10.500000 11.444444 10.111111 11.444444 10.888889
## [50] 11.055556 10.277778 10.500000 10.944444 10.777778 10.944444 10.444444
## [57] 11.055556 11.000000 11.055556 11.666667
The use of the if above is technically unnecessary, but it serves as a nice reminder that it only makes sense to take the difference between temperatures starting in year 2. Now with T_Celsius and Temp_ch calculated, we can make a nice table of years and temperature values and changes.
years <- 1912:1971;
dat <- cbind(years, nhtemp, T_Celsius, Temp_ch);
| years | nhtemp | T_Celsius | Temp_ch |
|---|---|---|---|
| 1912 | 49.9 | 9.944444 | NA |
| 1913 | 52.3 | 11.277778 | 1.33333333 |
| 1914 | 49.4 | 9.666667 | -1.61111111 |
| 1915 | 51.1 | 10.611111 | 0.94444444 |
| 1916 | 49.4 | 9.666667 | -0.94444444 |
| 1917 | 47.9 | 8.833333 | -0.83333333 |
| 1918 | 49.8 | 9.888889 | 1.05555556 |
| 1919 | 50.9 | 10.500000 | 0.61111111 |
| 1920 | 49.3 | 9.611111 | -0.88888889 |
| 1921 | 51.9 | 11.055556 | 1.44444444 |
| 1922 | 50.8 | 10.444444 | -0.61111111 |
| 1923 | 49.6 | 9.777778 | -0.66666667 |
| 1924 | 49.3 | 9.611111 | -0.16666667 |
| 1925 | 50.6 | 10.333333 | 0.72222222 |
| 1926 | 48.4 | 9.111111 | -1.22222222 |
| 1927 | 50.7 | 10.388889 | 1.27777778 |
| 1928 | 50.9 | 10.500000 | 0.11111111 |
| 1929 | 50.6 | 10.333333 | -0.16666667 |
| 1930 | 51.5 | 10.833333 | 0.50000000 |
| 1931 | 52.8 | 11.555556 | 0.72222222 |
| 1932 | 51.8 | 11.000000 | -0.55555556 |
| 1933 | 51.1 | 10.611111 | -0.38888889 |
| 1934 | 49.8 | 9.888889 | -0.72222222 |
| 1935 | 50.2 | 10.111111 | 0.22222222 |
| 1936 | 50.4 | 10.222222 | 0.11111111 |
| 1937 | 51.6 | 10.888889 | 0.66666667 |
| 1938 | 51.8 | 11.000000 | 0.11111111 |
| 1939 | 50.9 | 10.500000 | -0.50000000 |
| 1940 | 48.8 | 9.333333 | -1.16666667 |
| 1941 | 51.7 | 10.944444 | 1.61111111 |
| 1942 | 51.0 | 10.555556 | -0.38888889 |
| 1943 | 50.6 | 10.333333 | -0.22222222 |
| 1944 | 51.7 | 10.944444 | 0.61111111 |
| 1945 | 51.5 | 10.833333 | -0.11111111 |
| 1946 | 52.1 | 11.166667 | 0.33333333 |
| 1947 | 51.3 | 10.722222 | -0.44444444 |
| 1948 | 51.0 | 10.555556 | -0.16666667 |
| 1949 | 54.0 | 12.222222 | 1.66666667 |
| 1950 | 51.4 | 10.777778 | -1.44444444 |
| 1951 | 52.7 | 11.500000 | 0.72222222 |
| 1952 | 53.1 | 11.722222 | 0.22222222 |
| 1953 | 54.6 | 12.555556 | 0.83333333 |
| 1954 | 52.0 | 11.111111 | -1.44444444 |
| 1955 | 52.0 | 11.111111 | 0.00000000 |
| 1956 | 50.9 | 10.500000 | -0.61111111 |
| 1957 | 52.6 | 11.444444 | 0.94444444 |
| 1958 | 50.2 | 10.111111 | -1.33333333 |
| 1959 | 52.6 | 11.444444 | 1.33333333 |
| 1960 | 51.6 | 10.888889 | -0.55555556 |
| 1961 | 51.9 | 11.055556 | 0.16666667 |
| 1962 | 50.5 | 10.277778 | -0.77777778 |
| 1963 | 50.9 | 10.500000 | 0.22222222 |
| 1964 | 51.7 | 10.944444 | 0.44444444 |
| 1965 | 51.4 | 10.777778 | -0.16666667 |
| 1966 | 51.7 | 10.944444 | 0.16666667 |
| 1967 | 50.8 | 10.444444 | -0.50000000 |
| 1968 | 51.9 | 11.055556 | 0.61111111 |
| 1969 | 51.8 | 11.000000 | -0.05555556 |
| 1970 | 51.9 | 11.055556 | 0.05555556 |
| 1971 | 53.0 | 11.666667 | 0.61111111 |
In the next section, I will move on to consider a more complicated example using nested for loops (i.e., a for loop inside of another for loop).
The for loop in R: nested loops
Loops can be nested inside one another, such that the inner loop is run one time for each iteration of the outer loop. A common example of when nested loops are useful is in working with two dimensional arrays (e.g., tables or matrices). I will share a quick example from community ecology theory, in which species interactions within a community are often represented by square matrices like the one below,
Community ecology theory is not the focus here, so it is fine to skip a couple paragraphs to just move along to the coding problem. For more context though, each element in the above matrix defines how a slight increase in the density of one species affects the density of another species when species densities are at some equilibrium state. Each row and column in M represents a single species, so there are two species in the above matrix. Where rows and column numbers are identical, we have the diagonal of the matrix; this defines how a species affects its own density (i.e., self-regulation). The off-diagonals define how a slight increase in one species’ density affects a different species; in the above example, both species decrease each others densities because each has a negative affect on the other (the species in row 1 is negatively affected by species 2 by a magnitude of , and the species in row 2 is negatively affected by species 1 by a magnitude of ). If one of these two off-diagonal elements were positive and the other were negative (e.g., , ), we could interpret this as a predator-prey interaction. If both off-diagonal elements were positive (e.g., , ), we could interpret this as a mutualistic interaction.
To investigate community stability, theoreticians use random matrix theory to test how likely it is that communities with specific properties will return to equilibrium species densities when perturbed (e.g., Allesina and Tang 2015; Allesina and Tang 2012). Developing this theory sometimes requires generating many large M matrices with random interaction strengths (off-diagonal elements) but uniform interaction types (competitor, predator-prey, or mutualist) and self-regulation (diagonal elements). If we take the case of large M matrices in which all interactions are predator-prey (e.g., a big food web), all pairs of row-column elements need to have opposite signs. In other words, if is positive, then needs to be negative. To generate a random matrix with this property, we need go through the elements of M and change the signs of values where appropriate.
The coding issue is therefore to build a large matrix in which the sign of is the opposite of . We also want the absolute values of the off-diagonal elements to be random numbers, and the diagonal elements to be -1. These latter two properties can be made with the following lines of code, which will make a 10 ✕ 10 matrix as printed off below:
M_vals <- rnorm(n = 100, mean = 0, sd = 1);
M_vals <- round(M_vals, digits = 2);
M_mat <- matrix(data = M_vals, nrow = 10);
diag(M_mat) <- -1; # Adds -1 values to diagonal
print(M_mat);
## [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10]
## [1,] -1.00 -0.51 0.97 -0.62 -1.12 -1.82 -1.60 2.19 0.88 0.73
## [2,] 0.87 -1.00 0.04 0.23 0.03 -0.35 0.36 0.16 0.27 -0.31
## [3,] -0.28 1.27 -1.00 -0.19 -0.96 0.89 0.04 0.13 0.04 -1.07
## [4,] -0.66 -1.15 0.22 -1.00 0.78 0.47 -0.21 0.00 1.95 -0.42
## [5,] -1.08 1.78 -1.70 0.19 -1.00 0.22 -0.95 -1.60 -0.41 -1.65
## [6,] -0.53 -0.28 0.49 -0.50 1.75 -1.00 0.24 1.75 -0.18 -0.50
## [7,] -1.76 -0.74 2.12 1.12 -0.37 1.11 -1.00 -0.78 -1.24 -0.03
## [8,] -0.10 -1.55 -0.01 -0.67 -0.41 -0.25 -0.13 -1.00 0.35 0.85
## [9,] 0.88 2.45 0.92 1.39 -0.41 -0.51 -1.03 -0.63 -1.00 -0.45
## [10,] 0.31 -2.07 0.21 -1.05 -0.18 0.81 0.54 -0.89 0.69 -1.00
The above random matrix has diagonal elements all equal to -1, and off-diagonal elements independently drawn from a standard normal distribution
. The task is now to to make sure that pairs of off-diagonal elements
and
have opposite signs. In other words, if M_mat[1, 3] is positive, then M_mat[3, 1] should be negative (recall that R indices in brackets refer first to the row, then the column of a matrix: M_mat[row, column]). Unlike the previous problems in these notes, it is difficult to see how to create such a matrix without using loops (or editing the values by hand). We need to iterate over M_mat for each row and for each column, reversing the signs of off-diagonal elements whenever necessary. To do this, we can use a for loop within another for loop – the outer loop iterates over rows, and the inner loop iterates over columns. Whenever a pair of elements M_mat[i, j] and M_mat[j, i] are found to have the same sign, M_mat[i, j] is multiplied by -1.
N_species <- dim(M_mat)[1]; # Get total row & col number
for(i in 1:N_species){ # For each row in the matrix
for(j in 1:N_species){ # For each column in the row
if(i < j){ # Only need to look at upper triangle
elem_sign <- M_mat[i, j] * M_mat[j, i];
if(elem_sign > 0){
M_mat[i, j] <- -1 * M_mat[i, j];
}
}
} # Finish all columns in the row
} # Finish all rows
print(M_mat);
## [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10]
## [1,] -1.00 -0.51 0.97 0.62 1.12 1.82 1.60 2.19 -0.88 -0.73
## [2,] 0.87 -1.00 -0.04 0.23 -0.03 0.35 0.36 0.16 -0.27 0.31
## [3,] -0.28 1.27 -1.00 -0.19 0.96 -0.89 -0.04 0.13 -0.04 -1.07
## [4,] -0.66 -1.15 0.22 -1.00 -0.78 0.47 -0.21 0.00 -1.95 0.42
## [5,] -1.08 1.78 -1.70 0.19 -1.00 -0.22 0.95 1.60 0.41 1.65
## [6,] -0.53 -0.28 0.49 -0.50 1.75 -1.00 -0.24 1.75 0.18 -0.50
## [7,] -1.76 -0.74 2.12 1.12 -0.37 1.11 -1.00 0.78 1.24 -0.03
## [8,] -0.10 -1.55 -0.01 -0.67 -0.41 -0.25 -0.13 -1.00 0.35 0.85
## [9,] 0.88 2.45 0.92 1.39 -0.41 -0.51 -1.03 -0.63 -1.00 -0.45
## [10,] 0.31 -2.07 0.21 -1.05 -0.18 0.81 0.54 -0.89 0.69 -1.00
Note that in the matrix M_mat modified above, all pairs of off-diagonal elements M_mat[i, j] and M_mat[j, i] have opposite signs. Why did that work? We can start with the loops, the outer of which (for(i in 1:N_species)) started going through rows starting with row i = 1. While i = 1, the inner loop (for(j in 1:N_species)) went through all columns from 1 to 10 in row 1. Each unique combination of row i and column j identified a unique matrix element M_mat[i, j], and the code then checked to see if any action needed to be taken in two ways. First, the code checked to see if(i < j) – if not, then the whole bracketed if statement is skipped and we move on to the next column j. This if statement prevents the code from unnecessarily checking the same i and j pair twice, and prevents it from changing the diagonal where i == j. Second, the code assigning elem_sign checks to see if M_mat[i, j] and M_mat[j, i] have opposing signs by multiplying the two values together (two positives or two negatives multiplied together will equal a positive value for elem_sign; one positive and one negative will equal a negative value). If elem_sign > 0, then we know that M_mat[i, j] and M_mat[j, i] are either both positive or both negative, so we fix this by changing the sign of M_mat[i, j] (multiplying by -1). The figure below gives a visual representation of what is happening.
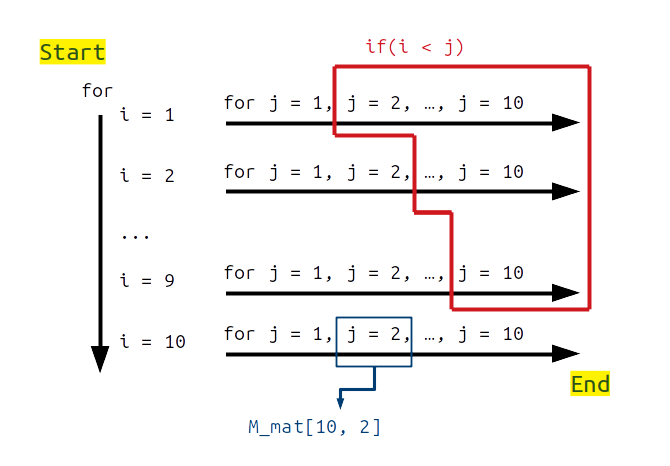
In the figure above, we start with the case in which i = 1 (i.e., the first row), and move through each value of j from j = 1 to j = 10. If M_mat[i, j] is in the upper right triangle of the matrix (i.e., i < j; shown in red), then the code checks to see if both M_mat[i, j] and M_mat[j, i] have the same sign. The loop ends when it has moved through all values of i from 1 to 10, hence looking at each element M_mat[i, j] in the matrix.
Once the logic of the nested loop makes sense, the rest comes down to remembering the syntax for for coding loops in R correctly. This comes with practice, so I have included some practice problems using loops below. Next, I will have a (more brief) look at the while loop in R. The general idea of iterating the same task many times will be the same, but the conditions under which the task is iterated will change slightly.
The while loop in R
The general idea of a while loop is the same as that of a for loop. In both cases, we are repeating the same task multiple times. But whereas we could specify the full range of values in which to substitute some value (e.g., i) within a for loop, in a while loop, we only specify the conditions under which to continue iterating. The printing of numbers from 1 to 10, as done with the for loop above, can also be done with the while loop below.
i <- 1;
while(i <= 10){
print(i);
i <- i + 1;
}
## [1] 1
## [1] 2
## [1] 3
## [1] 4
## [1] 5
## [1] 6
## [1] 7
## [1] 8
## [1] 9
## [1] 10
There are few important things to note.
- We need to specify an initial value of
i = 1(elseprint(i)will not return anything). - The loop will continue as long as
iis less than or equal to 10 (while(i <= 10)). - It is critical to increment
iwithin the loop (i.e., add a value of 1 at the end soi <- i + 1).
If we had forgotten number 3, then the value of i would always be 1. Aside from only printing the number 1 many times (rather than 1-10), notice that the loop would never actually terminate because i would always stay less than or equal to 10. This situation is called an ‘infinite loop’, and will result in a situation where it is necessary to terminate the loop manually (i.e., tell R to stop it, either using the red stop sign in the Rstudio console, or by holding down ‘CTRL + C’). This is almost always to be avoided, but to see what happens, remove the line i <- i + 1; and run the rest of code above.
Sometimes the use of for versus while loops is a matter of personal preference, such as with the simple example of printing a set of numbers from 1 to 10. In some cases, however, use of a while loop will make coding easier.
Consider a situation in which data need to be randomly sampled from a subset of 100 out of 1000 total different entities (the details do not matter – these entities could be different lochs, fields, or trees in a park). If we just need to sample 100 values out of 1000 without replacement, we can do this with a single line of R code:
subset <- sample(x = 1:1000, size = 100, replace = FALSE);
print(subset);
## [1] 561 672 746 389 941 106 606 571 193 519 510 629 951 33
## [15] 11 492 146 46 287 50 525 379 709 143 237 631 574 843
## [29] 732 687 274 236 591 328 508 856 388 751 90 599 220 392
## [43] 947 104 949 516 694 181 160 431 393 270 423 511 66 540
## [57] 842 649 892 972 446 272 71 877 244 817 22 907 612 676
## [71] 623 834 141 325 1000 495 635 665 715 769 749 398 307 222
## [85] 77 891 6 526 395 283 252 717 336 178 735 27 835 340
## [99] 821 474
This is easy enough, but what if, having already chosen these 100 entities, we decide that we need another 100, for a total of 200 unique samples (without replacement). We could find a creative way of using sample again in R (give this a try), but there is a logical way to do this with a while loop. The idea is to sample a single value from 1:1000, then check to see if that value is already in the subset. If it is in the subset, then throw it out and keep going. If it is not in the subset, add it. Continue until the size of subset is 200.
while(length(subset) <= 200){
samp <- sample(x = 1:1000, size = 1);
if(samp %in% subset == FALSE){
subset <- c(subset, samp);
}
}
print(subset);
## [1] 561 672 746 389 941 106 606 571 193 519 510 629 951 33
## [15] 11 492 146 46 287 50 525 379 709 143 237 631 574 843
## [29] 732 687 274 236 591 328 508 856 388 751 90 599 220 392
## [43] 947 104 949 516 694 181 160 431 393 270 423 511 66 540
## [57] 842 649 892 972 446 272 71 877 244 817 22 907 612 676
## [71] 623 834 141 325 1000 495 635 665 715 769 749 398 307 222
## [85] 77 891 6 526 395 283 252 717 336 178 735 27 835 340
## [99] 821 474 20 657 232 507 32 707 653 52 210 504 250 132
## [113] 439 805 537 592 95 595 358 730 411 208 505 70 555 322
## [127] 300 163 256 113 40 755 34 174 56 88 123 628 614 82
## [141] 363 61 601 651 893 875 743 733 138 437 114 895 897 345
## [155] 738 867 740 713 869 977 194 414 684 822 690 881 768 925
## [169] 500 622 277 227 991 221 128 166 689 989 910 859 823 418
## [183] 873 30 514 78 481 868 486 487 103 716 145 960 841 905
## [197] 69 133 675 701 731
The while loop above will continue as long as subset contains less than 200 numbers. If a randomly selected number from 1 to 1000 is not in the subset, then it is immediately added to make a bigger subset with the new number appended to it. The end result is that the above code has added 100 new unique values to the previous sample of 100.
Practice problems
Below are some practice problems for working with loops. To see the answers, click on the ‘Details’ arrows to the left at the bottom of each question. Note that your answers might differ from mine – there is more than one way to solve each problem!
- Using a
fororwhileloop, print all of the numbers from 1 to 1000 that are multiples of 17. (Hint: The mod operator%%returns the remainder after division. For example,14 %% 2would return a value of 2 because 14/4 = 3 with a remainder of 2.)
for(i in 1:1000){
if(i %% 17 == 0){
print(i);
}
}- In the
nhtemp, write a loop to add up the temperatures for all of the even numbered years, then divide by the total number of even numbered years to get the average.
Y <- 1912:1971; # Years
N <- length(nhtemp); # Total temps
A <- 0; # Added temp
C <- 0; # Count
for(i in 1:N){
if(Y[i] %% 2 == 0){
A <- A + nhtemp[i];
C <- C + 1;
}
}
avg_A <- A/C;
print(avg_A);- Using a
whileloop, calculate the sum of the series, to at least 10000 terms. What does the value appear to approach as more terms are added? (Hint: Useif(){}, or anif(){}else{}to switch from+to-)
val <- 0;
deno <- 1;
iter <- 1;
sign <- 1;
while(iter < 1000000){
if(sign < 0){
val <- val - (4/deno);
}
if(sign > 0){
val <- val + (4/deno);
}
sign <- -1 * sign;
deno <- deno + 2;
iter <- iter + 1;
}
print(val);- From here, write a while loop that prints out standard random normal numbers (use
rnorm()) but stops (breaks) if you get a number bigger than 1.
i <- 0;
while(i <= 1){
i <- rnorm(n = 1);
print(i);
}- Create an 8 ✕ 8 matrix
matwith diagonal values of 1 and off-diagonal values randomly selected from a standard normal distribution (usingrnorm()). Using nestedforloops as in the above notes, swap elementsmat[i, j]withmat[j, i]only ifmat[i, j] < mat[j, i](so that the higher number is in the lower triangle).
mat_v <- rnorm(n = 64, mean = 0, sd = 1);
mat_v <- round(mat_v, digits = 2);
mat <- matrix(data = mat_v, nrow = 8);
diag(mat) <- 1;
N <- dim(mat)[1];
for(i in 1:N){
for(j in 1:N){
if(mat[i, j] < mat[j, i]){
temp_val <- mat[i, j];
mat[i, j] <- mat[j, i];
mat[j, i] <- temp_val;
}
}
}
print(mat);Additional resources
- The Essence of Loops
References
Allesina, Stefano, and Si Tang. 2012. “Stability criteria for complex ecosystems.” Nature 483 (7388). Nature Publishing Group: 205–8. doi:10.1038/nature10832.
———. 2015. “The stability–complexity relationship at age 40: a random matrix perspective.” Population Ecology, 63–75. doi:10.1007/s10144-014-0471-0.
